Short Description
This segmentation method performs a N-class thresholding based on a K-Means classification of the image histogram, then extracts objects in a bottom-up manner using user-defined minimum and maximum object sizes. Very useful to detect clustered objects in fluorescence microscopy.
This book chapter proposes a video tutorial on the HK-Means plugin (see tutorial 2). Images used in the tutorial are available on this Zenodo repository.
Documentation
To cite this plugin, please refer to this paper: A. Dufour, V. Meas-Yedid, A. Grassart, and J.-C. Olivo-Marin, “Automated Quantification of Cell Endocytosis Using Active Contours and Wavelet”, Proc. ICPR 2008, Tampa, FL, USA.
Getting started
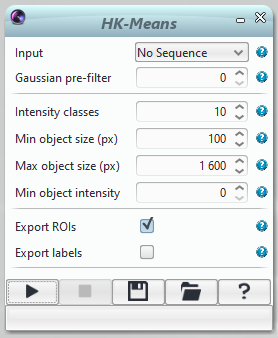
Input: Sequence to process.
Gaussian pre-filter: To pre-process the sequence with a Gaussian filter give an integer value >0 (sigma of the Gaussian blur in x,y for a 2D sequence and x,y,z for a 3D sequence)
Intensity classes: 10 by default. Increase this value if the objects of interest have different intensitye values.
Min object size (px): minimum size of clusters, in pixels. To estimate the size of objects (for instance nuclei) on a 2D sequence, draw a ROI around an object and check the value of its interior. To estimate object size on a 3D sequence, draw a 2D ROI on one slice (one Z position), and multiply the interior value of this ROI by the number of slices spanned by the object.
Max object size (px): maximum size of clusters, in pixels. See above how to estimate objects size.
Min object intensity: at least one pixel of the object must be of this intensity. Mouse over objects and background to evaluate the minimum objet intensity.
Export ROIs: outputs objects as region of interest overlayed on the original image
Export Labels: outputs a labelled image
Tracking
When working with a timelapse, you’ll see the option “Prepare for tracking”. If you tick it, ROIs detected by HK-Means will be sent to the Swimming Pool, a location that can be accessed by several plugins, including the Spot tracking plugin.
![]()
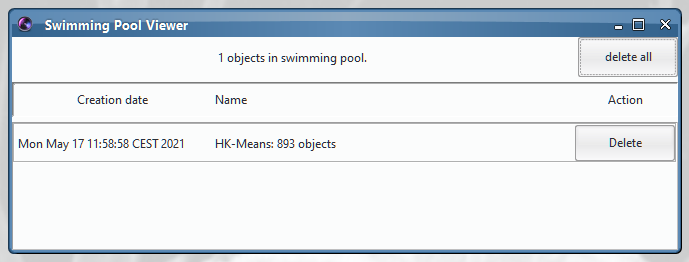
At any time, you can see the content of the Swimming Pool by opening the Swimming Pool viewer.
To further process your timelapse with the Spot Tracking plugin, click on “Select detection results here” to select the ROIs you just created with the HK-Means plugin. Then click on Run the Spot Detector plugin if you have a simple tracking case or go to Interface → Advanced interface to set advanced tracking options.
![]()
Block HK-Means
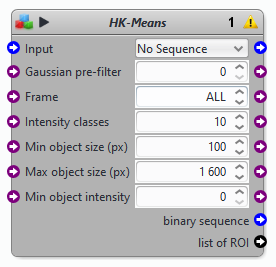
A few parameters in the block are different from the HK-Means plugin interface:
Frame: to apply HK-Means on all or only one frame.
binary sequence: output a binary sequence witth two labels (background and objects)
list of ROI: output objects as region of interest.
Note that, if the image has several channels and you would like to apply the HK-Means only on one channel, you need to use the block Extract channel first and input the extracted channel.
One review on “HK-Means”