Short Description
Characterizes the PSF in 3D in the entire volume of a light-sheet image containing fluorescent beads.
Related protocols: P1 – Detect tissue mimics and crop 3D sub-volume and P2 – SNR and contrast measurements. Related plugin: metroloSPIM. Tissue-mimics example datasets to test the plugin and protocols are available on Zenodo.
Team: R&D ImagerieInstitution: RESTORE-GOT IT Team UMR 5070-CNRS 1301-INSERM EFS Univ. Toulouse, France
Website: https://www.itav.fr/plateformes-technologiques/R-et-D-imagerie/Team: SLN - Team Loza
Institution: ICFO-Institut de Ciences Fotonique, Av. Carl Friedrich Gauss, 3, Castelldefels, 08860, Barcelona, Spain
Website: https://www.icfo.eu/lang/research/groups/groups-details?group_id=16Citation: https://doi.org/10.1038/srep44939
Documentation
This protocol was designed to characterize the PSF in 3D in the entire volume of a light-sheet image containing fluorescent beads.
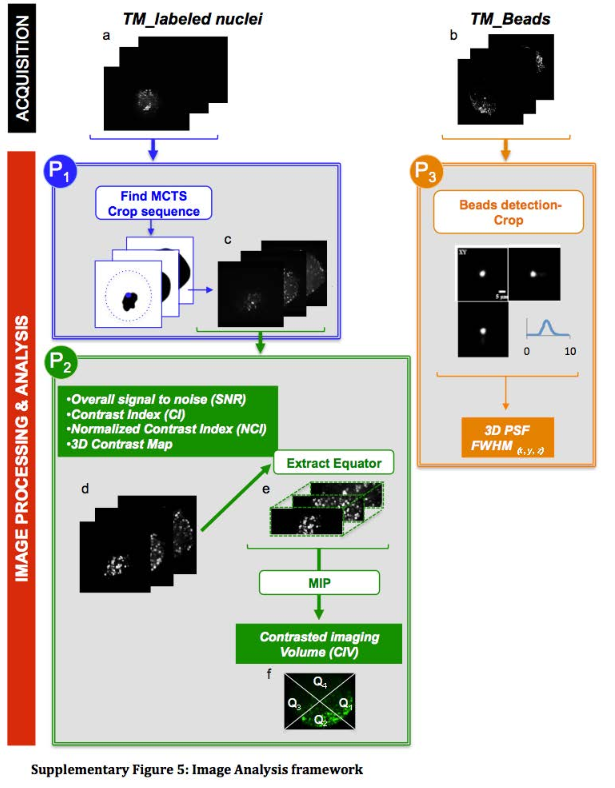
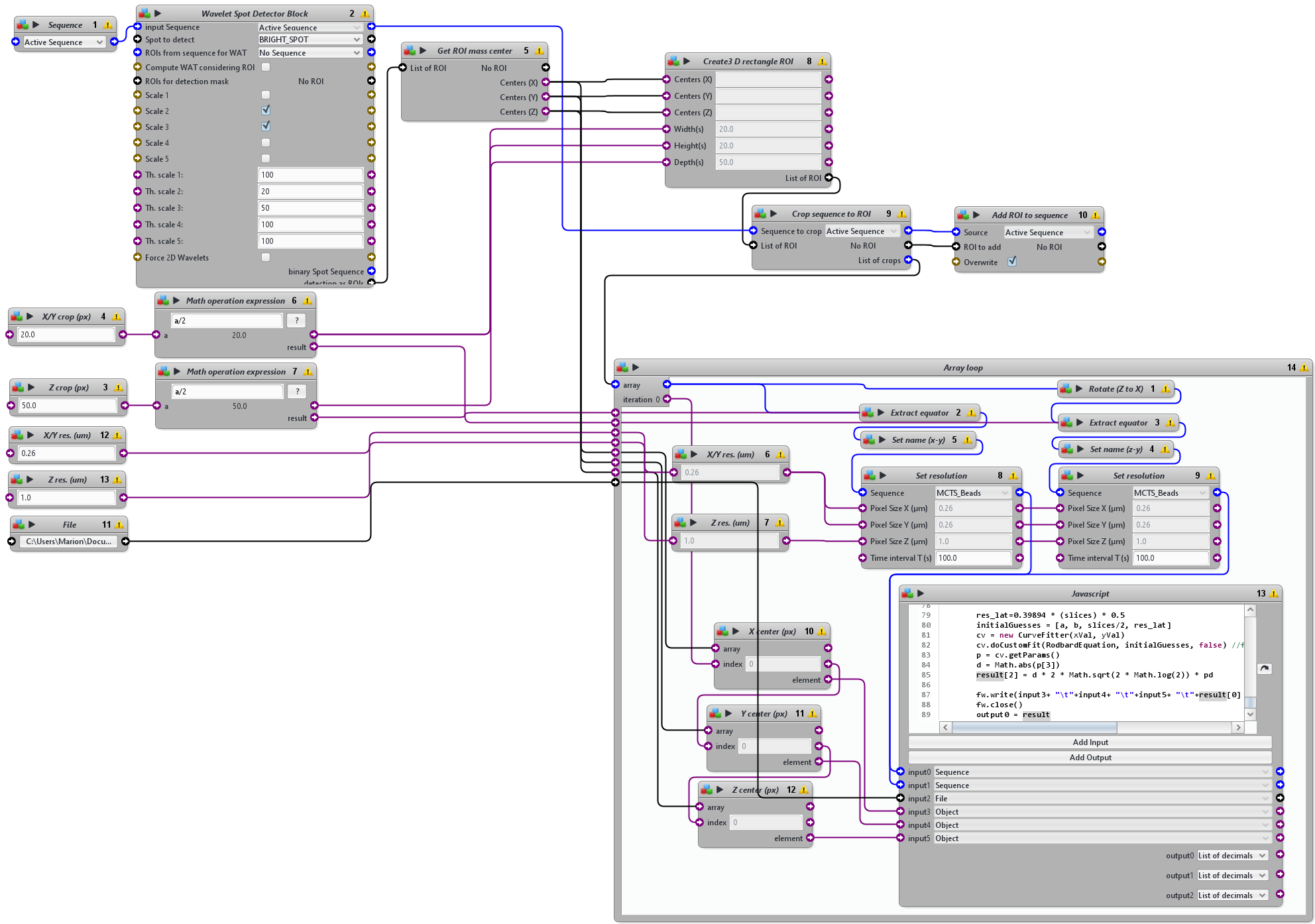
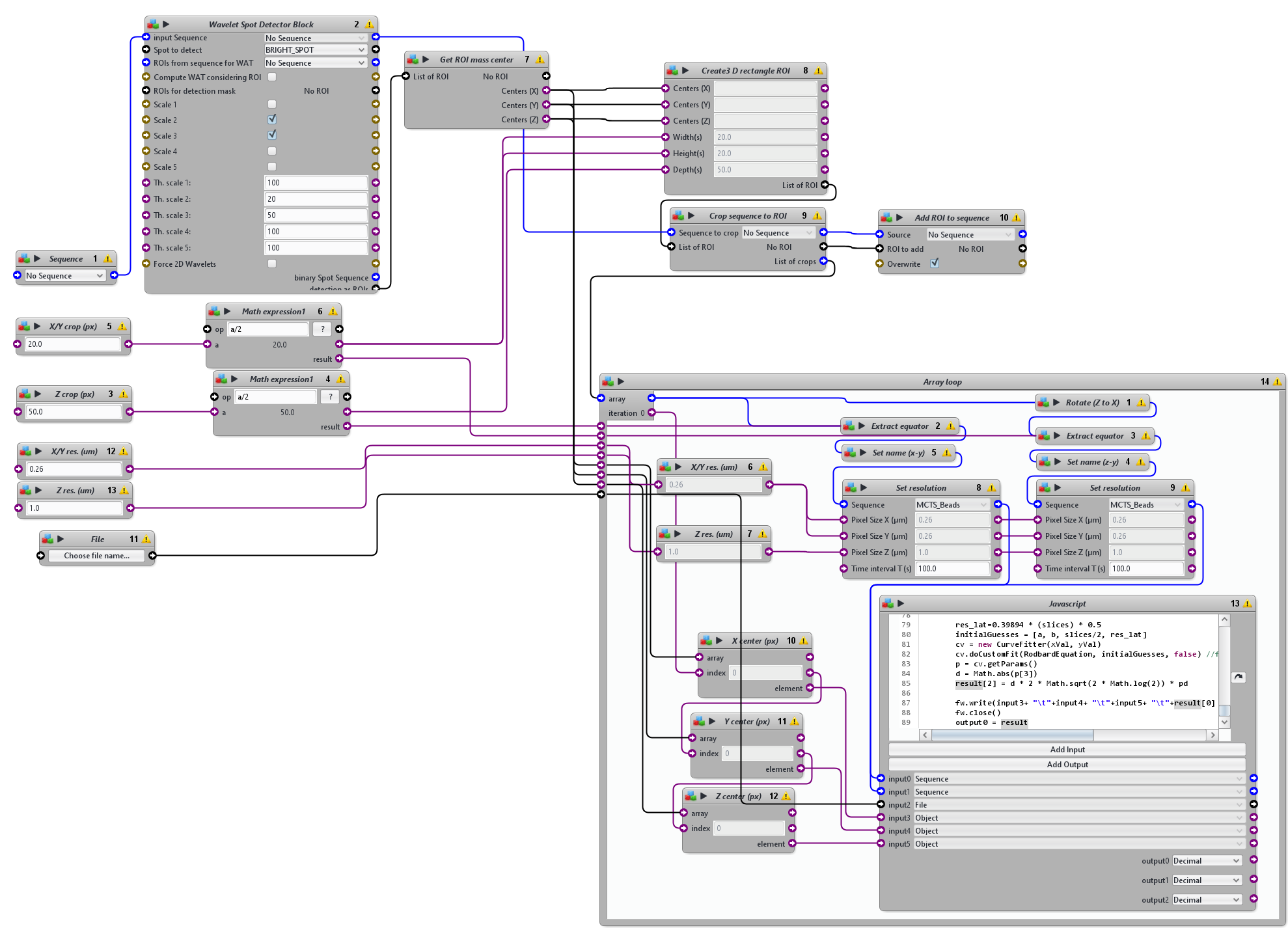
The protocol starts with a wavelet-based spot detection used to detect the positions of beads within the 3D volume and then extracts a sub-volume of six times the size of the expected 3D PSF. 2D Gaussian function is then fitted to the central profile of each sub-volume along the three axes(x, y, z) and the FWHM is then computed from the standard deviation of the Gaussian fit providing the actual size of the 3D PSF. The values of FWHM of the PSF in the entire volume in 3D are then listed in a text file.
Please cite Andilla et al. 2017 if you re-use this protocol or make a modified version of it. Figure below comes from the supplementals of Andilla et al., 2017.This protocol belongs to the bioimage analysis workflow described below. The protocol P1 detects tissue mimics imaged with a light-sheet microscope and crops 3D sub-volume, and the protocol P2 automatically measures the SNR, Contrast Index (CI), Normalized Contrast Index (NCI) and Contrasted Imaging Volume (CIV) on the extracted sub-volume.